Cell
All organism are composed of cells. Some of single(either Prokaryotic & Eukaryotic) & some of multi cells.
- Any thing less than a complete str. of cell doesn’t ensure independent living.
- Study of cell is called cytology.
- The word cell was first coined by Robert Hook in 1665.
Discovery of Cell & Its structure
The invention of the microscope & its improvement leading to the electron micro graph revealed all the strucural details of cell.
- Anton Von Leeuwenhoek first saw & discribed the live cell
- Robert Brown later discovered the nucleus.
- Theodore Schwan founded that all animal cell are covered by thin outer layer called Plasma Membrane & presence of cell wall is the unique character of plant cells.
Definition of the Cell
- Cell is the smallest enclosed chemical system of open thermodynamic nature possessing a unique structural cohesion & showing functional self sustainability, reproducibitity, irritability & adaptiveness.
- Cell represents life at its smallest level -hence it is also called the str, functional & reproductive units of life.
The significance of the cell in this way was first put forward by the cell theory.
Cell Theory or Cell Doctrine
It states that
- all organism are composed of similar units of organisation, called cells, and also that
- all cell arise from the preexisting cells.
In 1838-39, Theodor Schwann ( a Zoologist) & Matthias Schleiden (a Botanist) contributed the earliest version of the cell theory.
- The cell is the unit of str. , physiology & organisation in living organism.
- The cell retains the dual existence as a distinct entity & a building block in the construction of the organism
- Cells form by free-cell formation, similar to the formation of crystals (spontaneous generation)
We know today that third point is wrong.
In 1855, Rudolph Virchow put forwarded the correct interpretation of cell formation by cell division
- Rudolph Virchow’s dictum,” Omnis cellula e cellula”, which means that all cells arise from pre-existing cells.
Exception to cell Theory
- Viruses (Alive inside host yet are not made up of cells)
- The first cell did not originate from a pre-existing cell.
Introduction to the Cell
- Cells come in different sizes and shapes and perform different activities
- Mycoplasmas, the smallest cells, are only 0.3 μm in length while bacteria could be 3 to 5 μm
- Among multicellular organisms, human red blood cells are about 7.0 μm in diameter.
- Longest cell in human body is Neuron
- Biggest cell is egg of Ostrich.
- The biggest single celled organism is Acetabularia.
Cell are of two types : Prokaryotic & Eukaryotic cells
Prokaryotic cells

- Animals having these cells are called Prokaryotes & basically placed in Monera kingdom & include members from two most primitive domains of life i.e Bacteria & Archean . These are the simplest type of cellular organisms.
- Characteristic feature
- They are characterized by absence of well defined nucleus or have pro-nucleus. They genetic material is naked & c/a Nucleoid(Nuclear body)
- In addition to genomic or main DNA, many bacteria have small circular DNA Plasmids.
- They don’t have membrane bounded organells.
- Represented by bacteria, blue green algae, mycoplasma etc.
- Generally smaller & multiply more rapidly than the eukaryotic.
- Vary greatly in shape & size & ususly around 0.5 μm to 10 μm of diameter.
- All unicellular, either solitary or colonial
- Multiplication is by binary fission.
- Cell Envelop
3 layered str.
- Glycocalyx ( Some have capsule or some have slime layer form)
- Capsule
- Some bacteria form a thick outer capsule of high molecular weight, viscous polysaccharid gel
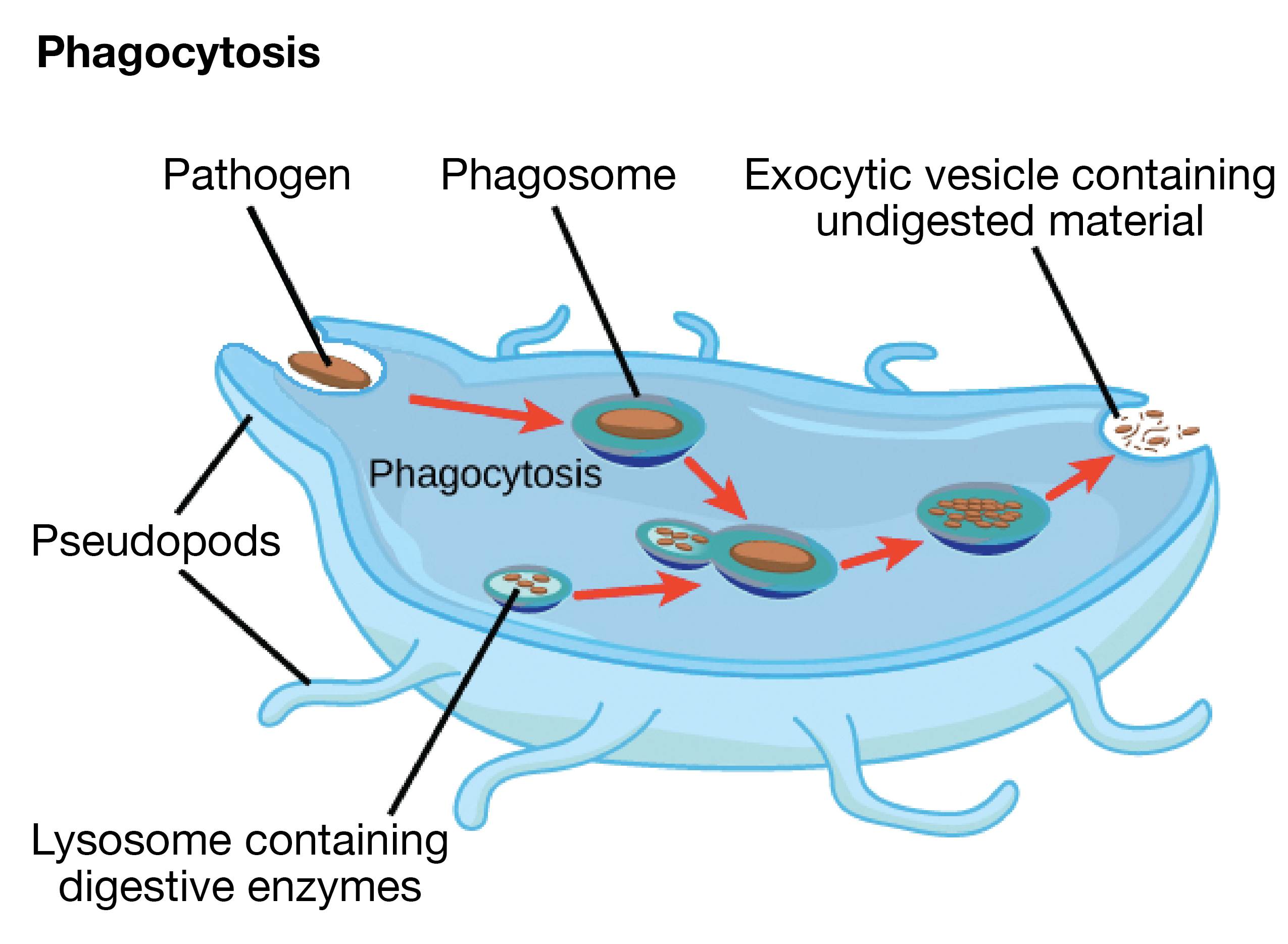
- capsules confer resistance to phagocytosis.
- Slime Layer
- Glycocalyx in the form in loss sheath i.e amorphous
- Capsule
- Cell wall
- It is a rigid str. made up of peptidoglycans,
- Peptidoglycans are unique to prokaryotic organisms
- These walls also contain Teichoic Acids & Lipoteichoic Acids.
- confer the characteristic cell shape
- Provide mechanical protection.
- Two groups of bacteria lakes CW
- Mycoplasma species
- The L form
- that arises form either Gram +ve or -ve bacteria cells & have lost their ability to produce the peptidoglycan str.
- The outer membrane of Gram – ve bactria is composed of lipopolysaccharides.
- It is a rigid str. made up of peptidoglycans,
- Plasma Membrane
- semipermeable in nature ; interacts with outside world
- composed primarily of protein & phospholipid (about 3:1)
- perform many functions, including transport, biosynthesis & energy transduction.
- str. similar to the eukaryotes.
- Mesosome
- special memberane str. formed by extension of plasma memberan into the cell.
- extension are in form of
- vesicles
- tubules
- lamellae
- helps in
- cell wall formation
- DNA replication & distribution to daughter cells
- Respiration
- Secretion processes,
- To increase the surface area of the plasma membrane & enzymatic content.
- Chromatophores
- Memeranous extension in some cynobacteria
- externsion in cytoplasm contains pigments.
- Staining
- Bacteria can be classified into two groups on the basis of the differences in the cell envelops & the manner in which they respond to the staining procedure developed by Gram {Gram Stain – }

- Gram +ve
- take up the Gram stain
- Gram -ve
- don’t take up the gram stain.
- Gram +ve
- Bacteria can be classified into two groups on the basis of the differences in the cell envelops & the manner in which they respond to the staining procedure developed by Gram {Gram Stain – }
- Surface Structures
- Flagella
- differ in str. from eukaryotic flagella.
- Composed of 3 parts
- Filament (longest portion)
- Hook
- Basal body
- Basal body anchored in PM & CW gives rise to cylindrical filament.
- moves by whirling about its long axis.
- the no. & arrangement of flagella – are diagnostically useful.
- Pilli
- Pili are slender, hairlike, proteinaceous appendages on the surface of many bacteria.( spl. Gram -ve)
- imp in adhesion to host surface or rocks in streams.
- Fimbriae
- small bristle like fibres sprouting out of the cell.
- imp in adhesion to host surface or rocks in streams.
- Flagella
- Cytoplasmic Structures
- Ribosome
- cytoplasm is densely packed with 70S ribosomes.(made up of 50S+30S)
- these are associated with plasma membrane
- They are 15nm-20nm in size
- are the site of protein synthesis.
- Polysome
- Several ribosomes attach to a single mRNA to form
- ribosomes of polysome translate the mRNA into proteins.
- Inclusion Bodies
- other granules represent metabolic reserves (e.g poly β -hydroxybutyrate, polysaccharide, polymetaphosphate, and meta chromatic granules.
- These are not bounded by any membrane system
- lie free in cytoplasm
- Gas Vacuoles
- found in blue green & purple & green photosynthetic bacteria.
- Endospores
- Bacillus & Clostridium species can produce endospores
- heat resistant, resting cells that are formed intracellularly & contain a genome & all essential metabolic machiery.
- The endospore is protected in a complex protective spore coat.
- Ribosome
- Nucleoid
- bacterial nucleoid, which contains the DNA fibrils, lacks a limiting membrane
- The bacterial nucleoid is a str. contrain a single circular DNA without any histone protein.
- It is about 4-8 Mbp large & with a molecular weight of appox. 3×109. The unfold DNA is about 1mm long.
| Type of Cells |
Eukaryotic cell
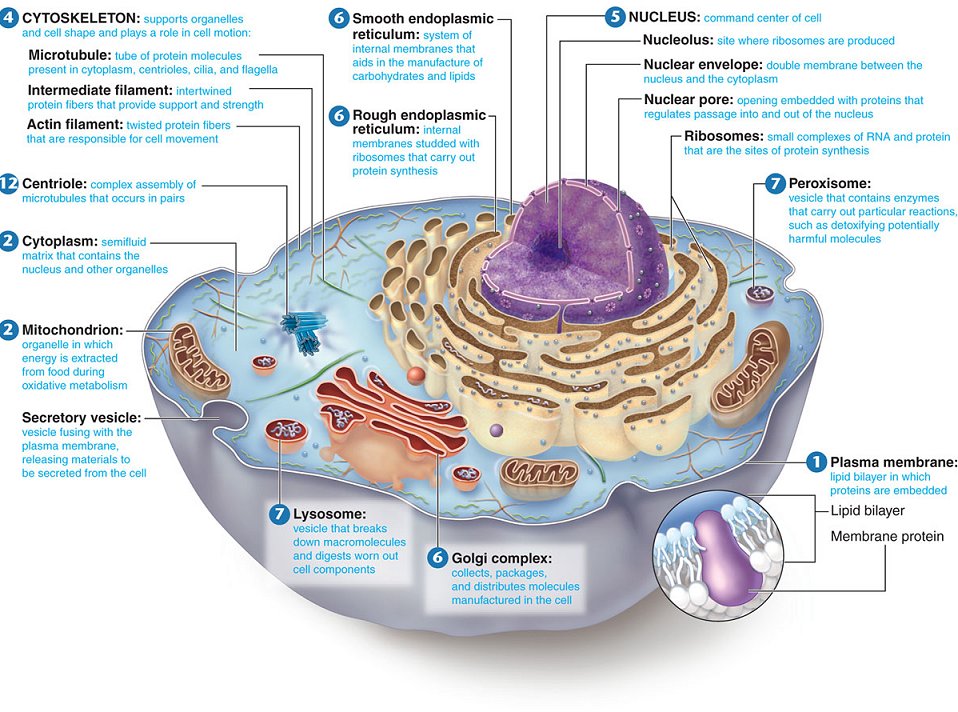
- Include all the protists, plants,animals & fungi.
- In eukarotic cells there is extensive compartmentalization of cytoplasm through presence of membrane bounded organelles.
- posses organised nucleus with nuclear envelope.
- Genetic material is organised into chromosome.
- these have variety of complex locomotory & cytoskeletal str.
Structure of Typical cell
- Cell Wall
- in plant & fungi there is a rigid cell wall
- Non living & freely permeable.
- 1° wall is capable of growth, which diminishes as the cell matures
- 2° wall is formed on the inner side of the cell.
- Made up of cellulose, hemicellulose, pectine, proteins (Plants) or Chitin(Fungi)
- Function
- Provide shape & rigidity to cell.
- Protect from mechanical damage & infection
- it helps in cell to cell interaction
- provides barrier to undesirable macromolecules.
- Middle Lamella
- layer mainly of calcium pectate
- holds or glues the different neighboring cell togather
- Plasmodesmata
- cell wall & middle lamellae transverded by plasmodesmata
- connet the cytoplasm of neighboring cells.
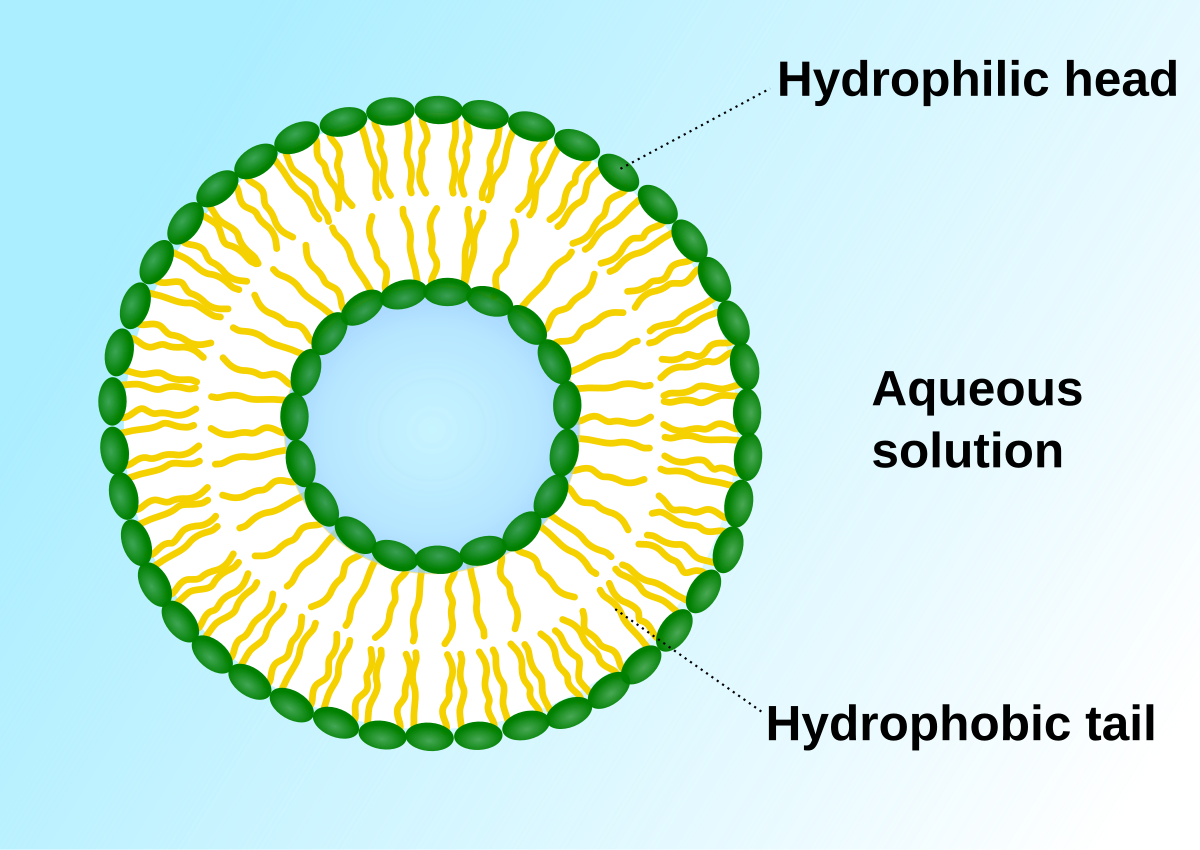
- Cell Membrane
- also k/n as Plasma membrane. & form the outer covering of the animal cell, in plant cell found within cell wall.
- composed of lipids & proteins mainly, also contain choesterol & Charbohydartes ; found by the chemical studies on the RBC
- Fluid Mosic Model given by Singer & Nicolson,1972 to explain the str.
- Major lipids are arranged in the membrane with
- polar head toward the outer sides \
- Hyddrophobic tails towards inner part ; so that is it remain protected from aq. environment.
- Major lipids are arranged in the membrane with
- Lipids are in quasi fluid nature so allow lateral movement, which is important for many functions like cell growth, formatn fo intercellular juctions, secretion,endocytosis, cell division etc
- Function
- Transport of chemical across it. it is selective permeable.
- Protoplasm
The whole fluid present inside plasma membrane is k/n as protoplasm.
Name given by Purkenje in 1839
Made 80 % water
Inorganic : Organic :: 81:19
Compostion
- Oxygen – 76 %
- Carbon – 10.5 %
- Hydrogen – 10 %
- Nitrogen – 2.5 %
- Cytoplasm : Fluid outside the nuclear membrane
- Nucleoplasm : fluid inside the nuclear membrane
- Endomemrane System

each membranous organells is distinct in tems of str. & fun. but considered ES b/c functions are cordinated.
It regulates protein traffic & performs metabolic function in the cell
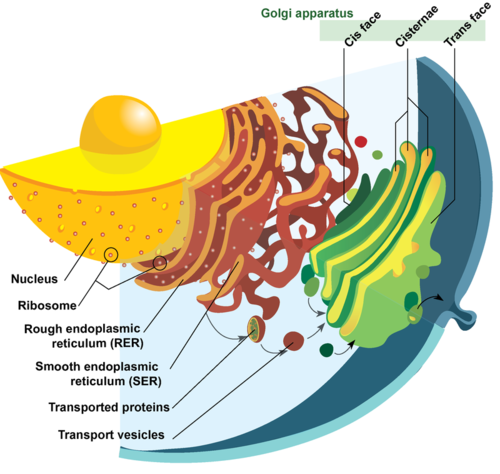
- Endoplasmic Reticulum

- extensive network of membrane bound tubules & sacs
- Membrane separates lumen from cytosol
- continuous with the nuclear envelope
- Functions of Smooth ER (not having ribosomes on surface)
- Synthesis of lipids
- in animal cells lipid like steroidal hormones
- metabolism of carbohydrates
- Ca2+ storage
- detoxification of drugs & poisons
- Functions of Rough ER (having ribosomes on surface) – extensive & continuous with outer membrane of nucleus.
- Aids in sythesis of secretory & other proteins form bound ribosomes ;
- adds carbohydrates to glycoproteins
- produces new membrane
- Golgi Apparatus
- Camillo Golgi (1898) first observed densely stained reticular str. , made up of tubes, vesicles & vacuoles.
- Stacks of flattened membranous sacs (cisternae) ; has polarity (cis & trans faces)
- Cis (Convex or forming face) & traces (Concave trans or the maturing face) of organelle are entirely different but interconnected.
- Function
- Packaging material
- material to be packed came from ER in form of Vesicles fuse with the cis face
- then move towards the maturing face
- delivered either to the intra cellular targets or secreted outside the cell.
- material to be packed came from ER in form of Vesicles fuse with the cis face
- modification of proteins synthesised by ribosomes in the cisternae of GA.
- Important site of formation of glycoproteins & glycolipids
- synthesis of many polysaccharides
- sorting of Golgi products, which are then released in vesicles
- Packaging material
- Lysosome
- Membrane bound vesicular str. of hydrolytic enzymes (in animal cells) formed by process of packaging in the GA
- hydrolytic enzymes, optimally active at acidic pH.
- Suicidal bags of the cell
- Function
- These enzymes are capable of digesting carbohydrates (carbohydrases), proteins (proteases), lipids (lipases) & nucleic acids.
- breakdown of ingested substances, cell macromolecules & damaged organelles for recycling.
- Vacuole
- large membrane bounded vesicles in plants , can occupy 90 % vol of plant cell.
- bounded by single membrane c/a tonoplast.
- Function
- digestion, storage, waste disposal, water balance, cell growth & protection.
- tonoplast facilitates ions & materials against conc. gradients into the vacuole.
- Contratile vacuole : imp. for osmoregulation & excreation in Amoeba
- Food Vacuoles : in protists these formed by engulfing food particles.
Mitochondria & Chloroplasts change energy from one form to another.
- Mitochondrion

- bounded by double membrane & no. varries depending on the physiological activity of cell.
- inner compartment is c/a matrix.
- Inner membrane has infoldings c/a Cristae
- toward the matrix
- to increase the surface area
- they have a single circular DNA, few RNA molecules, ribosomes(70S) & components for synthesis of proteins.
- These divide by fission.
- In case of human, maternal mitochondria in the baby.
- Function
- site cellular respiration
- produce cellular enery in the form of ATP so c/a Power house of cell
- bounded by double membrane & no. varries depending on the physiological activity of cell.
- Plastids

- found in all plant cells & euglenoides
- bear specific pigments & impart colors to plants. based on that
- Chloroplast
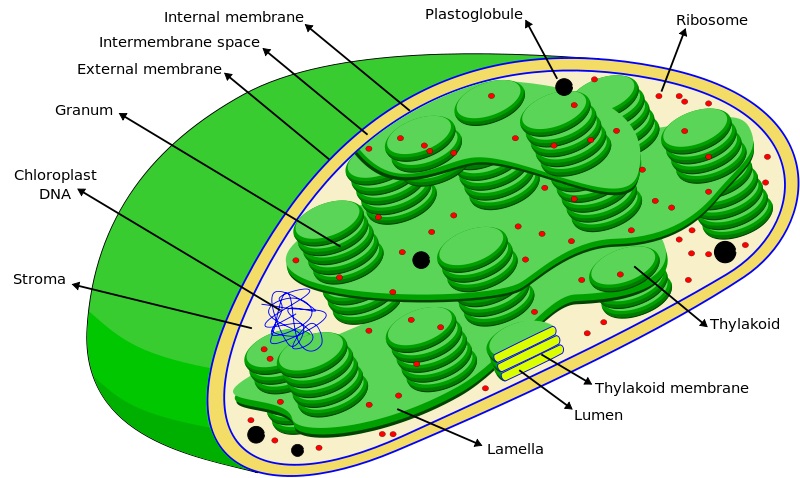
- typically two membranes around fuid stroma, which contains membranous thylakoids stacked into grana (In plants)
- stroma lamellae connect thylakoid of different grana.
- lens shaped, oval, spherical,discoid or even ribbon like organelles having variable length & width
- have chlorophyll II & carotenoid pigments in thylakoids
- stroma of contains enzymes required for synthesis of carbohydrates & proteins. & double stranded circular DNA & ribosomes(70S)
- majority of these are found in mesophyll cells of leaves.
- Function
- Photosynthesis : help in trapping light energy required for photosynthesis
- Chromoplasts
- containss fat soluble carotenoid pigments like carotene, xanthophylls & others are presents
- They give part of plant yellow, orange or red colour
- Leucoplasts
- are colourless plastides or varied shapes & size with stored nutrients
- Function
- Store nutrients
- Amyloplasts – carbohydrates (Starch)
- Elaiplasts – oils & fats
- Aleuroplasts – Proteins
- Store nutrients
- Peroxisome
- specialized metabolic compartment bounded by a single membrane
- Function
- contains enzymes that transfer hydrogen to water, producing hydrogen peroxide (H2O2) as a by product, which is converted to water by other enzymes in the peroxisome
- Cytoskeleton
- filamentous proteinaceous str. in cytoplasm
- Function
- Mechanical support
- Mobility
- Maintenance of the shape of cell
- Flagella
- differ in str. from prokaryotic flagella.
- Composed of 3 parts
- Filament (longest portion)
- Hook
- Basal body
- Basal body anchored in PM & CW gives rise to cylindrical filament.
- moves by whirling about its long axis.
- the no. & arrangement of flagella – are diagnostically useful.
- Cilia
- small str. work like oars, causing the movement of either cell or the surrounding fluid.
- Centrosome & Centriole
- centrosome is an organelle usually containing 2 cylindrical str. called centrioles.
- Cylindrical structures respnosible for spindle fibre formation which help in cell division
- They are also basal bodies in cilia and flagella
- Microbodies
- membrane bound minute vesicles , contain various enzymes
- present in both plant & animal cells.
The eukaryotic cell’s genetic instructions are housed in the nucleus & carried out by the ribosomes
- Ribosomes
- granular str. first observed under e– microsope as dense particel by George Palade (1953)
- Two subunits made of ribosomal RNA & proteins ; \
- 80 S → 60 S + 40 S
- 70 S → 50 S + 30 S
- S stands for Svedberg’s Unit : sedimentation coefficient →indirectly a measure of density & size.
- can be free in cytosol or bound to ER
- Functions
- Protein synthesis
- Nucleus
- first described by Robert Brown in 1831.
- Flemming stained with basic dyes c/a Chromatin
- In a cell which is not dividing (interphase cell) has highly dense nucleoprotein fibers called chromatin
- Has two layers of nuclear membranes surrounding it
- Membrane has nuclear pores
- The nuclear matrix also has a spherical nucleolus – site for active RNA synthesis
- Actively dividing cells, structured chromosomes are present instead of nucleus
- A single human cell has around 2 meters of DNA distributed among its 46 chromosomes
- usually the easiest organelle to see under a microscope.
All eukaryotic cells are not identical.
- Plant cells
- Animal cells
DNA The Genetic Material
- Deoxyribonucleic Acid : It is the genetic material of the cell
- DNA is +nt in the chromosomes which are located in the nucleus of the cell
- amount of DNA per unit cell ∝ No. of chromosome per unit cell.
- DNA as acidic material +nt in nucleus first identified by Friedrich Meischer in 1869. & named it Nuclein.
- Evidence of DNA is genetic material
- Indirect
- Most of the DNA is located in chromosome i.e is in nucleus where as protein & RNA are also abundant in cytoplasm
- molecular composition of DNA in all cell is same
- diploid cell contain twice amount of DNA than haploid
- Direct
- Frederick Griffith in 1928
- O.T. Avery, C.M. Macleod & M. McCarty in 1944
- A.D Hershey & M. Chase in 1952(Unequivocal proof) – with bacteriophage virus & E.Coli – Radioactive Phosphorus
- Indirect
Structure of DNA
- Nucleic Acids (DNA & RNA) are macromolecules composed of repeating subunits called Nucleotides
- Each nucleotide consists of
- A phosphate group
- A five-carbon sugar (Pentose)
- In DNA – 2‐deoxyribose (thus the name deoxyribonucleic acid);
- in RNA, – ribose (thus ribonucleic acid)
- Nitrogen containing compound called a base :
- In DNA ; adenine, guanine, thymine, and cytosine
- In RNA in place of thymine there is Uracil
- Adenine & Guanine – double ring bases c/a Purines
- Cytosine, thymine & Uracil – Single ring bases c/a Pyrimidines
- The nitrogenous base is linked to the pentose sugar through a N-glcosidic linkage to form Nucleoside.
- when a phosphate group is linked to 5′-OH of nucleoside through phosphoester linkage, a corresponding nucleotide (or deoxynucleotide) is formed.
- two nucleotide linked through 3′-5′ phosphodiester linkage to form a dinucleotide. more nucleotide join to form polynucleotide chain.
- The polymer thus formed
- the back bone of polynucleotide chain is due to sugar & phosphates
- free phosphate moiety at 5′-emd of ribose sugar – 5′ end of polynucleotide chain
- free 3′-OH group referred as 3′-end of polynucleotide chain.
- The N bases linked to sugar moiety project from the backbone.
- the back bone of polynucleotide chain is due to sugar & phosphates
- In 1953, James Watson & Francis Crick,based on X ray diffraction data produced by Maurice Wilkins & Rosalind Franklin, proposed Double Helix model of str. of DNA.
- They proposed that DNA exists as a double helix in which the two polynucleotide chains are coiled about one another in a spiral
- The two polynucleotide strands are held together in their helical configuration by hydrogen bonding between bases in opposing strands
- The resulting base‐pairs are stacked between the two chains perpendicular to the axis of the molecule like the steps of a spiral staircase.
- base paring confers unique property to the polynucleotide chain. i.e complementary nature (not identical)
- If the sequence of bases in one strand of DNA double helix is known, the sequence of bases in the other strand is also known because of the specific base‐pairing.
- The sugar‐phosphate backbones of the two complementary strands are antiparallel; that is they have opposite chemical polarity.
- This is represented by 3’ End & 5’ End
- The bond is always b/w
- Adenine & Thyamine with 2 H bonds
- Guanine & Cytosine with 3 H bonds
- Analogous hydrogen bonding between cytosine and adenine, for example, is not possible except when they exist in their rare structural states (when mutations occur).
- The base pairs in DNA are stacked 3.4 nm w
- one of the hollmarks of their of their proposition was base pairing b/w two strands of polynucleotide chains.
- This was based on the observation of Erwin Chargaff that for a dounble stranded DNA, the ratio b/w A & T & G&C are constant & equal to 1.
- A+G = T+C
- A+T= specific for species.
C+G
DNA Replication
- Mechanism of DNA replication is proposed by Watson & Crick
- It takes place in the S phase of cell division
- The precision of the DNA replication is the function of nitrogenous bases, N bases are the only variable component of DNA is the nitrogenous base.
- The four different nitrogenous bases, however, vary in sequence and quantity from organism to organism and from species to species – A, C, G, T
- There is always of consistency of C+T = A+G
- consistency is a function of
- the size of the bases,
- purines are larger molecules than pyrimidines.
- the space available between the sugar molecules in the double helix, and
- Within the diameter of the DNA helix, only purine‐ pyrimidine paring can occur;
- purine‐purine would be too large to fit into the helix an
- pyrimidine‐ pyrimidine pairing would be too small.
- the chemical nature of these bases
- the size of the bases,
- consistency is a function of
Process of DNA Replication
The scheme suggested that the two strands would separate & act as a template for the synthesis of new complementary strands.
After the completion of replication, each DNA molecule would have one parental & one newly synthesised strand.
This scheme was termed as semiconservative DNA replication.
- Origin of Replication (Ori)
- There is definite region in DNA where the replication originates these are c/a Origin of replication.
- Prokaryotes have single origin of replication but Eukaryotes has thousands.
- Activation of Nucleotide
- deoxyribonuleodie monophosphate → deoxyribonuleodie triphosphate
- activared nucleotide serve dual function
- acting as substrates
- provide energy for polymerization of rxn
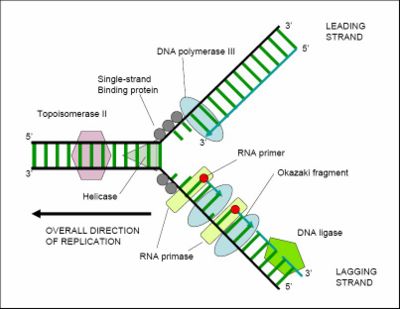
- Unwinding of Helix by Helicase (ATP dependent)
- For long DNA molecule, since the two strands can’t be separated in its entire length (due to very high energy requirement)
- The replication occur within a small opening of the DNA helix referred to as replication fork.
- Single stand of replication fork are stabilized by SSBP
- Due to Unwinding supercoling occur. This tension is released by topoisomerase & in prokaryotes by DNAgyrase.
- Formation of Primer Strend
- DNA polymerase III is unable to initiate DNA synthesis as it is unable to deposit first nucleotide at the ori site.
- Primase synthesis short RNA primar. Once initiation start it enzymetically it is removed.
- Elongation by DNA Polymerase
- The DNA dependent DNA polymerase catalyse polymerisation only in one direction i.e 5’→3′.
- consequently on one strand of replication fork it is continuous, while on the other template with polarity 5’→3′ , it is discontinuous.
- The discontinuously synthesised fragments c/a Okazaki Fragments, are later joined by the enzyme DNA ligase.
- From the nucleoplasm, new molecules of the bases are bonded to the exposed bases.
- Because of the specificity of base‐pairing, the original partners of the exposed bases are replaced by new but chemically identical molecules.
Significance of DNA Replication
- all living org. perpertuate their kind through reproduction
- Reproduction entails a faithful transmission of genetic information of the parents to the progeny
- Since the DNA is the storehouse of genetic information, replication of DNA is the essence of reproduction and transmission of genetic traits
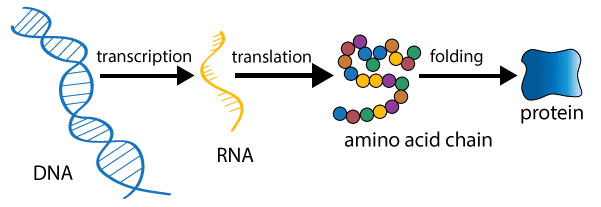
Central Dogma
Francis crick proposed the Central dogma. It state that the genetic information flow from DNA→RNA→Protein

Reverse Central Dogma or Teminism
found by H.Temin & D. Baltimore
RNA→DNA
Transcription
- The process of coping genetic information from one strand of the DNA into RNA is termed as transcription.
- The principle of complementarity governs the process of transcription

- A transcription unit in DNA
- A Promoter – at 5′ end of the str. gene referred in reference to the coding strand.
- The structural gene
- Template strand – polarity 3’→5′. act as template for transcription
- Coding strand – don’t code but strangle called coding strand.
- A terminator -at 3′ end of the str. gene referred in reference to the coding
- DNA dependent RNA polymerase also catalyse the polymerisation in only one direction. i.e 5’→3′.
- RNA polymerase I – rRNA
- RNA polymerase II – heterogeneous nuclear RNA (hnRNA) – precurosr of mRNA
- RNA polymerase III – tRNA, 5srRNA & snRNAs(Small nuclear RNAs)
- Types of RNA & the process of Transcription

- Initiation
- occurs when the enzyme RNA polymerase binds to a region of a gene called the promoter
- This signals the DNA to unwind so the enzyme can ‘‘read’’ the bases in one of the DNA strands.
- The enzyme is now ready to make a strand of mRNA with a complementary sequence of bases.
- Elongation
- Elongation is the addition of nucleotides to the mRNA strand.
- RNA polymerase reads the unwound DNA strand and builds the mRNA molecule, using complementary base pairs
- Termination
- occurs when RNA polymerase crosses a stop (termination) sequence in the gene.
- The Pre mRNA or hnRNA strand is complete, and it detaches from DNA.
- Initiation
Processing of hnRNA or Pre mRNA

- Splicing –
- The area of DNA which is transcribed but is excised is called the intron and the sequence retained is exon
- intron are removed & exons are jointed in a defined order.
- Capping – methyl guanosine triphosphate is added to the 5’end of hnRNA
- Tailing – adenylate(Poly-A) residues (200-300) are added at 3′ end in a template in independent manner.
Once the mRNA leaves nucleus and enters cytoplasm, it binds to the ribosomes
mRNA gets attached to several ribosomes called polysomes or polyribosomes
Transfer RNA

- In cytoplasm, there’s another type of special RNA that participates in protein synthesis called tRNA,Also referred to as sRNA or Soluble RNA
- tRNA has segments which are both double stranded and single stranded
- Double strands are not different as in DNA but areas where different parts of tRNA molecule fold over and base pair
- Result is a folded structure that resembles a clover leaf
- One end of the tRNA has the sequence CCA
- It is joined at the CCA end by a single molecule of an amino acid
- The sequence of amino acids is determined by triplet codon on mRNA
- At the apex of the one of the loops is a sequence of bases called anticodon which is the area of attraction between tRNA & mRNA
- The anticodon is complementary to the sequence of bases on mRNA
- Other two loops are place of tRNA’s attachment to ribosomes
Ribosome

- Ribosomes are made of proteins and rRNA
- mRNA gets attached to several ribosomes called polysomes or polyribosomes
Protein
- Proteins are essential quality of macromolecules that contribute to the structure and function of the cell
- Enzymes – that determine metabolic activities
- One Gene One Enzyme Hypothesis (1941) – Beadle & Tatum
- Genes control chemical reactions by regulating or determining the specificities of enzymes
- Before them, Archibald Garrod (1901) – Alkaptonuria – Inborn Errors of Metabolism – linked biochemical defects to genes
- PKU – Phenylketonuria (Phenylalanine breakdown missing)
- All tree RNAs are needed to synthesise a protein in a cell.
- mRNA – provides the template
- tRNA – brings aa & reads the genetic code.
- rRNA -play str. & catalytic role during translation.
Translation
- Protein synthesis occurs in the cytoplasm of the cell
- the mRNA carry the genetic message from the gene to the ribosome, the organelle associated with protein synthesis in the cytoplasm of cells.
Steps of Translation


- Transcription & Processing of mRNA
- As soon as the mRNA is formed, it leaves the nucleus and reaches the cytoplasm where it attaches with the ribosomes.
- Attachment of mRNA to ribosome
- The ribosome is uniquely designed to bring mRNA and tRNA together.
- The mRNA is threaded through the small subunit tunnel, while the large and subunit fit together to form tRNA binding pockets.
- mRNA gets attached to several ribosomes called polysomes or polyribosomes
- Activation of Aminoacidic
- amino acids occur in the cytoplasm in an inactive state.
- Each amino acid is activated by a specific activating enzyme known as amino acyl synthetase and ATP.

- Charging of tRNA
- The enzyme is recognised by the enzyme site of the tRNA and the amino acid residue is transferred to the amino acid attachment site of the tRNA.
- The amino acid is linked to the 3-OH at the -CCA end of tRNA. As a result, the AMP and enzyme are released and the final product amino acyl-tRNA is formed.

- Initiation of Protein Synthesis
- The mRNA usually has AUG as its first codon. AUG codes for methionine.
- The methionine is formulated and plays an important role in initiating the process of protein synthesis.
- After the protein synthesis is completed, the formyl methionine is detached by the activity of the hydrolytic enzyme.
- Elongation of Polypeptide chain
- whole process involves arrival of tRNA-amino acid complex, peptide bond formation and translocation
- amino acids are linked to one another
- A charged tRNA with the anticodon complementary to the codon of mRNA is brought to the A site of the ribosome.
- Peptide formation occurs between the amino acids in the P site and the A site in the presence of the enzyme, peptidyl transferase.
- The next step of elongation is the movement of ribosomes by one codon along the mRNA. This is called translocation.
- Termination & Release of Polypeptide Chain
- At the end of the mRNA chain there is a stop or terminator codon. There are three terminator codons – UAA, UGA and UAG.
- When the information processing region encounters a terminator codon, it signals the stop of protein synthesis.
- The ribosome separate from the mRNA chain and the ribosome dissociates into two subunits.
- Modification of Released Polypeptides
- The released polypeptide is in its primary form.
- It undergoes coiling and acquires a secondary and tertiary structure.
- It may combine with other polypeptides to assume a quaternary structure.
- Proteins known as chaperones help in the folding of the proteins to become functional.
Proteins synthesised on free ribosomes are released into the cytoplasm and function as structural and enzymatic proteins. Proteins formed on attached ribosomes, pass through the tunnel into the channels of ER and are exported as cell secretions by exocytosis after packaging in the Golgi apparatus.
Other Aspects of Protein Synthesis
- Most mRNAs are short-lived – destroyed once their messages are translated
- It is necessary to waste them as their presence may result in continuous translation of a particular protein disturbing the balance of the cell and are dangerous – Ex. Adrenalin !!**#@!!
- If a protein is continuously needed, then more mRNAs are continuously synthesized – feedback from cytoplasm to nucleoplasm
- Some mRNAs for proteins like Hemoglobin are stable as this is stored in abundant quantity in RBCs
- tRNAs and ribosomes are much stable & take part in multiple functions
Genetic Code
- The messages for a particular protein must be encoded in nitrogenous bases as the sugar and phosphate are essentially same in all cells and all DNA
- There are 20 Amino Acids; and there are 4 Bases in DNA
- Can the code be a double code?
- Then there will be only 16 combinations like A‐T, A‐C, A‐ G, A‐A, C‐T, C‐C, C‐G, G‐C etc. (remember we are talking about sequence of bases on one strand of DNA)
- Whatever the code is, it has to allow for at least 20 different combinations for 20 different amino acids
- The code has to be a Triplet Code as this accounts for 64 combinations
- Scientists discovered the code is a triplet code and have interpreted all of them
- Ex. UUU – Phenylalanine; AGU – Serine etc.
- Each triplet representing an amino acid is a codon
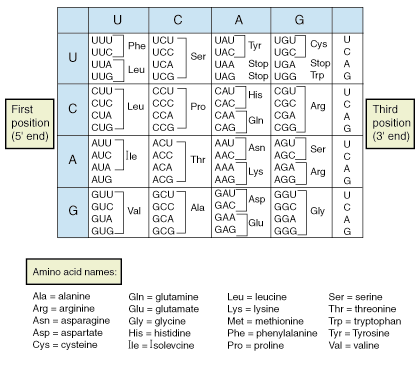
- 64 possible combinations are more than enough for 20 amino acids
- There is a certain redundancy in genetic code as more than one type of triplet codon exists for a amino acid
- It has been found by Francis Crick that only the first two bases are important for codon and the third base can be any of the four and the codon still translates into the same amino acid – This is called Wobble Hypothesis
Gene Mutations
- In any species, variations can be caused
- Changes in environment
- Changes in heredity constitution
- More commonly by interaction between both
- If any variation appears when environment changes and is restored when the environment is restored, then it is not inherited – Ex. Muscles – Exercise
- These are modifications – phenotypic differences between organisms with similar genotype
- Mutations are changes in the genetic makeup of an individual, which can be qualitative or quantitative
- Mutations are random in location
- The effects of mutations can be positive, negative or neutral
- Mutations can be classified variously according to
- Type of cells
- Size and quality
- Origin
| Somatic Mutation | Gametic Mutation |
| • Occur in body cells / non-reproductive cells Genetic and evolutionary consequences are insignificant, since only single cells or their daughter cells are involved However, if they occur during embryonic stage, impact is significant as large proportion of cells is involved • Ex. Some cancerous growths | In sperms and ova Heritable and immense genetic significanceThey are the raw material for natural selection |
| Size & Quality | |
| Point Mutation | Gross Mutation |
| Occur in very small segment of DNA, a single nucleotide or a pairCan be classified into | Involve a greater portion of a gene or entire geneCan occur due to rearrangement of genes within genome |
| Origin | |
| Spontaneous Mutation | Induced Mutation |
| Occur naturally / randomly in nature and their origin is unknown | Certain environmental conditions can induce mutationsAgents that can induce mutations – MutagensRadiations – X Rays, Gamma Rays, Ultraviolet raysTemperatureChemicals – Some Alkoloids, Benzene, Sodium Azide |
Packaging of DNA Helix
- DNA is slightly -ve charged molecule.
- In eukaryotic cells, there is set of +ve charged basic(due to abundance of lysines & arginines aa residue) protein c/a histones.
- Histon organised to form a unit of 8 molecule c/a histone octamer.
- The -ve charged DNA molecule molecule warp around it to form a str. c/a nucleosome. contain 200bp of DNA helix.
- Nucleosome constitute the repeating unit of a str. in nucleus c/a chromatin, thread like stained (coloured) bodies seen in nucleus.
- A nucleosomes in chromatin are seen as beads on string str. when viewed under electron microscope.
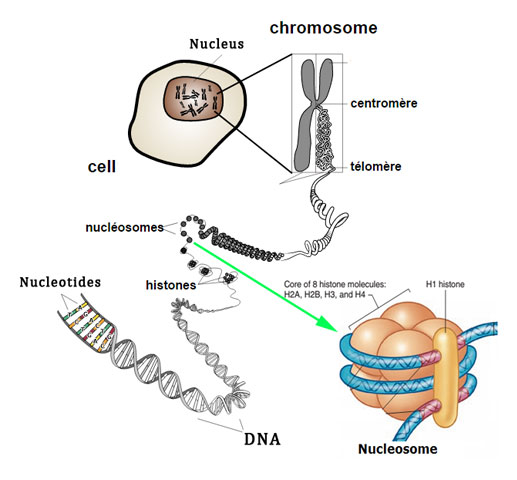
- these packed to form chromatin fivers that are further coiled & condensed at metaphase stage of cell division to form chromosomes.
- The packaging of chromatin at higher level requires additional set of proteins that collectively are referred to as Non histone chromosomal (NHC) proteins.
- Euchromatin – transcriptional active chromatin
- Hetrochromatin – in active
Chromosome
- are complex str. located in the cell nucleus visible only in dividing cells.
- interphase nucleus has a loose and indistinct network of nucleoprotein fibres called chromatin
- To get the cell with their chromosomes in this condensed state they are exposed to a mitotic inhibitor which blocks formation of the spindle & arrests cell division at metaphase stage.
- Some ex. of tissues used to obtain chromosome peripheral blood, bone marrow, amniotic fluid, and products of conception.
- Study of chromosomes is Karyology
Chromosomal Str.
- Composed of DNA, histone and non-histone proteins, RNA & polysaccarides. They are basically the ” packages” that contain the DNA.
- Under microscope they appear as thin, thread like str.
- Every chromosome essentially has a primary constriction or the centromere on the side of which disc shaped str. called kinetochores are present.
- Centromere holds the two chromatids of a chromosome.
Types of Chromosome
Based on the position of the centromere, chromosome can be classified into four types
- Metacentric Chromosome – middle centromere forming two equal arms of the chromosome.
- Sub-metacentric Chromosome – centromere slightly away from the middle resulting one shorter & one longer arm
- Acrocentric Chromosome – centromere situated close to its end forming one extremely short & one very long arm.
- Acrocentrics frequently have a secondary constrictions connecting small pieces of DNA called stalks & satellites
- Telocentric Chromosome – These have terminal centromere.
Sometimes a few chromosomes have non-staining 2° constriction at a constant location. They gives the appearance of a small fragment c/a Satellite.
Ideogram
- The ideogram is basically a “chromosome map” showing the relationship between the short and long arms, centromere (cen), and in the case of acrocentric chromosomes the stalks (st) and satellites (sa).
- The specific banding patterns are also illustrated.
- Each band is numbered to aid in describing rearrangements.
Some info about Human Chromosome
- Normal human somatic cells have 46 chromosomes: diploid
- 22 pairs, or homologs, of autosomes (chromosomes 1‐22) and
- two sex chromosomes
- Females carry two X chromosomes (44,XX) while
- males have an X and a Y (44,XY).
- Germ cells (egg and sperm) have 23 chromosomes:
- one copy of each autosome plus a single sex chromosome –
- This is referred to as the haploid number.
- One chromosome from each autosomal pair plus one sex chromosome is inherited from each parent
- Mothers can contribute only an X chromosome to their children while fathers can contribute either an X or a Y.
- Chromosome 1 is the designation for the largest human chromosome.
Cell Division
It is the process by which a mature cell divides and forms two nearly equal daughter cells which resemble the parental cell in a number of characters.
- Discovery: Prevost and Dumas (1824) first to study cell division during the cleavage of zygote of frog.
- Nagelli (1846) was the first to propose that new cells are formed by the division of pre-existing cells.
- Rudolf virchow (1859) proposed “omnis cellula e cellula” and “cell lineage theory”.
- A cell divides when it has grown to a certain maximum size which disturb the karyoplasmic index (KI)/Nucleoplasmic ratio (NP)/Kernplasm connection.
- Two processes take place during cell reproduction.
- Cell growth: (Period of synthesis and duplication of various components of cell).
- Cell division: (Mature cell divides into two cells).
- Cell cycle: Howard and Pelc (1953) first time described it. The sequence of events which occur during cell growth and cell division are collectively called cell cycle. Cell cycle completes in two steps:
- Interphase
- period between the end of one cell division to the beginning of next cell division.
- also c/a resting phase or not dividing phase.But, it is actually highly metabolic active phase, in which cell prepares itself for next cell division.
- In case of human beings it will take approx 25 hours.
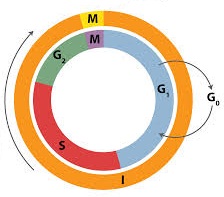
- Interphase is completed in to three successive stages.
- G1 phase/Post mitotic/Pre-DNA synthetic phase/Gap Ist
- S-phase/Synthetic phase
- G2-phase/Pre mitotic/Post synthetic phase/gap-IInd
- M-phase/Dividing phase/Mitotic Phase
- Nuclear division i.e. karyokinesis
- occurs in 4 phases
- prophase,
- it is an extended state in Meiosis I & further divided
- Leptotene
- Zygotene
- Pachytene
- Diplotene
- Diakinesis
- it is an extended state in Meiosis I & further divided
- metaphase,
- anaphase and
- telophase.
- prophase,
- It takes 5-10% (shortest phase) time of whole division.
- occurs in 4 phases
- Cytokinesis :
- Division of cytoplasm into 2 equal parts.
- In animal cell, it takes place by cell furrow method and
- in plant cells by cell plate method.
- Nuclear division i.e. karyokinesis
- Interphase
- Duration of cell cycle: It depends on the type of cell and external factors such as temperature, food and oxygen. Time period for G1, S, G2 and M-phase is species specific under specific environmental conditions.
- Regulation of cell cycle: Stage of regulation of cell cycle is G1 phase during which a cell may follow one of the three options.
- It may start a new cycle, enter the S-phase and finally divide.
- It may be arrested at a specific point of G1 phase.
- It may stop division and enter G0 quiscent stage. But when conditions change, cell in G0 phase can resume the growth and reenter the G1 phase.
- Cell division is of three types, Amitosis, Mitosis and Meiosis.
- Meiosis I is followed by a short-lived and transient interkinesis. There is no DNA replication at this stage.
- Meiosis II is more like mitosis and each haploid cell forms 2 more haploid cells so, after both meiotic divisions, we get 4 haploid cells from one diploid cell.
- Difference between cell Mitosis and Meiosis
| Characters | Mitosis | Meiosis |
| I. General | ||
| Site of occurrence | Somatic cells and during the multiplicative phase of gametogenesis in germ cells. | Reproductive germ cells of gonads. |
| Period of occurrence | Throughout life. | During sexual reproduction. |
| Nature of cells | Haploid or diploid. | Always diploid. |
| Number of divisions | Parental cell divides once. | Parent cell divides twice. |
| Number of daughter cells | Two. | Four. |
| Nature of daughter cells | Genetically similar to parental cell. Amount of DNA and chromosome number is same as in parental cell. | Genetically different from parental cell. Amount of DNA and chromosome number is half to that of parent cell. |
| II. Prophase | ||
| Duration | Shorter (of a few hours) and simple. | Prophase-I is very long (may be in days or months or years) and complex. |
| Subphases | Formed of 3 subphases : early-prophase, mid-prophase and late-prophase. | Prophase-I is formed of 5 subphases: leptotene,zygotene,pachytene, diplotene anddiakinesis. |
| Bouquet stage | Absent. | Present in leptotene stage. Chromosomes starts condendsing in leptotene |
| Synapsis Characterized by synaptonemal complex. | Absent. | Pairing of homologous chromosomes in zygotene stage.chromosomes appear as bivalent or tetrad. |
| Chiasma formation and crossing over.thus formation of recombination nodule. | Absent. | Occurs during pachytene stage of prophase-I. enzyme responsible for crossing over is Recombinase |
| Disolution of synaptoneamal complex & Chaismata formation | absent | In Diplotene stage :-Synaptonemal complex dissolve and homologous chromosomes separate from each other, except at the crossovers forming Chiasmata (the ‘X’ shaped structure).human primary oocytes remain in this stage until puberty when ovulation occurs Lampbrush chromosomes found in the oocyte of amphibians are formed at the diplotene stage |
| Disappearance of nucleolus and nuclear membrane | Comparatively in earlier part. | Comparatively in later part of prophase-I. i.e Diakinesis. also chiasmata separate. In prophase II , the nuclear envelope disappears. |
| Nature of coiling | Plectonemic. | Paranemic. |
| Theoretical | Chromosomes untangle and condenseTwo chromatids attached to the centromere can be seen clearlyEach of the duplicated centrosomes radiates microtubules (asters)Mitotic apparatus constitutes spindle fibres and astersGolgi bodies, nucleolus, endoplasmic reticulum and nuclear membrane disappear | |
| III. Metaphase | ||
| Metaphase plates | Only one equatorial plate | Two plates in metaphase-I : but one plate in metaphase-II. |
| Position of centromeres | Lie at the equator. Arms are generally directed towards the poles. | Lie equidistant from equator and towards poles in metaphase-I while lie at the equator in metaphase-II. |
| Number of chromosomal fibres | Two chromosomal fibre join at centromere. | Single in metaphase-I while two in metaphase-II. |
| Theoretical | Complete disintegration of the nuclear envelopeTwo sister chromatids attached by the centromere aligned at the equator, i.e. metaphase plateEach chromatid is attached to spindle fibres from opposite poles at kinetochores | Metaphase IBivalent chromosomes align at the equator and homologous chromosomes get attached to the spindles from opposite poles |
| IV. Anaphase | ||
| Nature of separating chromosomes | Daughter chromosomes (chromatids with independent centromeres) separate. | Homologous chromosomes separate in anaphase-I while chromatids separate in anaphase in anaphase-II. |
| Splitting of centromeres and development of inter-zonal fibres | Occurs in anaphase. | No splitting of centromeres. Inter-zonal fibres are developed in metaphase-I. |
| Theoretical | Splitting of centromere and two sister chromatids separate and go towards the opposite polesSister chromatids now become the daughter chromosomes | Homologous chromosomes move to opposite poles. Unlike mitosis, sister chromatids remain attached |
| V. Telophase : | ||
| Occurrence | Always occurs | Telophase-I may be absent but telophase-II is always present. |
| Theoretical | Chromosomes cluster at opposite poles and decondenseNuclear envelope develops around each cluster of chromosomes and two daughter nuclei are formedThe nucleolus, endoplasmic reticulum and Golgi apparatus are reformed | Telophase I :Nucleolus & Nuclear envelope reappears, chromosome collect at polesTelophase II :Nuclear membrane reappears |
| VI. Cytokinesis | ||
| Occurrence | Always occurs | Cytokinesis-I may be absent but cytokinesis-II is always present. |
| Nature of daughter cells | 2N amount of DNA than 4N amount of DNA in parental cell. | 1 N amount of DNA than 4 N amount of DNA in parental cell. |
| Fate of daughter cells | Divide again after interphase. | Do not divide and act as gametes. |
| Process | separation of cytoplasm takes place after two nuclei formed. cell organelles get distributed b/w daughter cells. | Cytokinessis I :dyad of haploid cells are formedCytokinessis :we get 4 haploid daughter cells, i.e. tetrad |
| VII. Significance | ||
| Functions | Helps in growth, healing, repair and multiplication of somatic cells.Occurs in both asexually and sexually reproducing organisms. | Produces gametes which help in sexual reproduction. |
| Variations | Variations are not produced as it keeps quality and quantity of genes same. | Produces variations due to crossing over and chance arrangement of bivalents at metaphase-I. |
| In evolution | No role in evolution. | It plays an important role in speciation and evolution. |
- Types of Mitosis
- Anastral mitosis: It is found in plants in which spindle has no aster.
- Amphiastral mitosis: It is found in animals in which spindle has two asters, one at each pole of the spindle. Spindle is barrel-like.
- Intranuclear or Promitosis: In this nuclear membrane is not lost and spindle is formed inside the nuclear membrane e.g. Protozoans (Amoeba) and yeast. It is so as centriole is present within the nucleus.
- Extranuclear or Eumitosis: In this nuclear membrane is lost and spindle is formed outside nuclear membrane e.g. in plants and animals.
- Endomitosis: Chromosomes and their DNA duplicate but fail to separate which lead to polyploidy e.g. in liver of man, both diploid (2N) and polyploid cells (4N) have been reported. It is also called endoduplication and endopolyploidy.
- Dinomitosis: In which nuclear envelope persists and microtubular spindle is not formed. During movement the chromosomes are attached with nuclear membrane.
- Types of meiosis: On the basis of time and place, meiosis is of three types
- Gametic/Terminal meiosis: In many protozoans, all animals and some lower plants; meiosis takes place before fertilization during the formation of gametes. Such meiosis is described as gametic or terminal.
- Zygotic or Initial Meiosis: In fungi, certain protozoan groups, and some algae fertilization is immediately followed by meiosis in the zygote, and the resulting adult organisms are haploid. Such a meiosis is said to be zygotic or initial. This type of life cycle with haploid adult and zygotic meiosis is termed the haplontic cycle.
- Sporogenetic Meiosis
(a) Diploid sporocytes or spore mother cells of sporophytic plant, undergo meiosis to form the haploid spores in the sporangia.
(b) Haploid spore germinates to form haploid gametophyte which produces the haploid gametes by mitosis.
(c) Haploid gametes fuse to form diploid zygote which develops into diploid sporophyte by mitotic divisions. e.g. in higher plants like pteridophytes, gymnosperms and angiosperms.
Mitosis
Meosis I
Meosis II